Project Overview Page¶
Clicking on the title of a project on the project list or project pull-down list at the top of the page will bring you to that project’s overview page. Important: Only users listed as researchers on the project may access this page. Example project overview page:
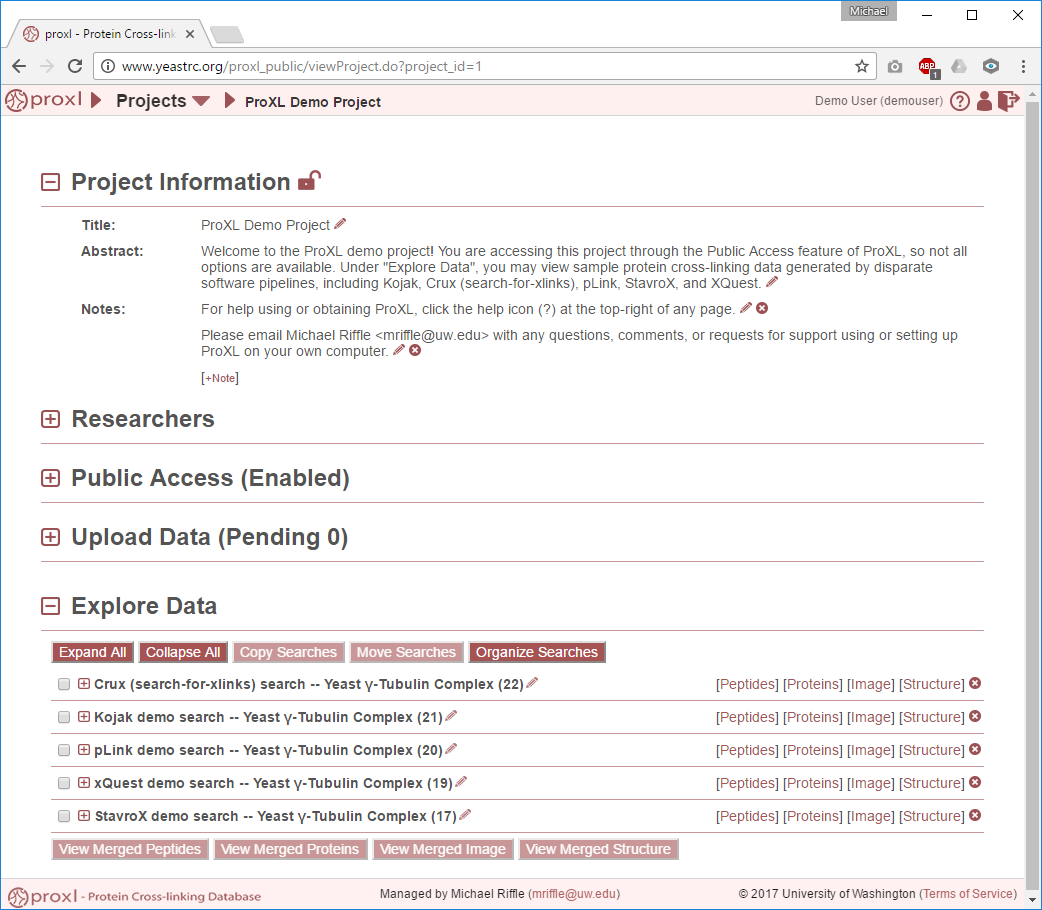
From here, you may edit the project information, add or remove researchers, toggle public access, explore or manage the data, and lock the project. Click the [+] icon next to each section to expand and view that section. Please see below for help on all of the sections.
Project Information¶
Project members may edit the title or abstract by clicking the pencil icon next to the respective item. Project members may add notes to the project by clicking “[+Note]”. Notes may be edited or deleted by clicking on the associated pencil or delete (X) icons. The title, abstract, and notes are visible to public users (if public access is enabled, see below).
Lock Project¶
The project may be locked by clicking the lock icon next to the “Project Information” section header:
Clicking the icon changes the icon to a locked state:
Projects that are locked may not be changed in any way until they are unlocked. This includes associated users, public access, uploading or annotating data, and so on. This is meant to accompany projects that are supporting publications, and so should not change. To unlock a project, the project owner may click on the lock icon next to the “Project Information” section header.
Researchers¶
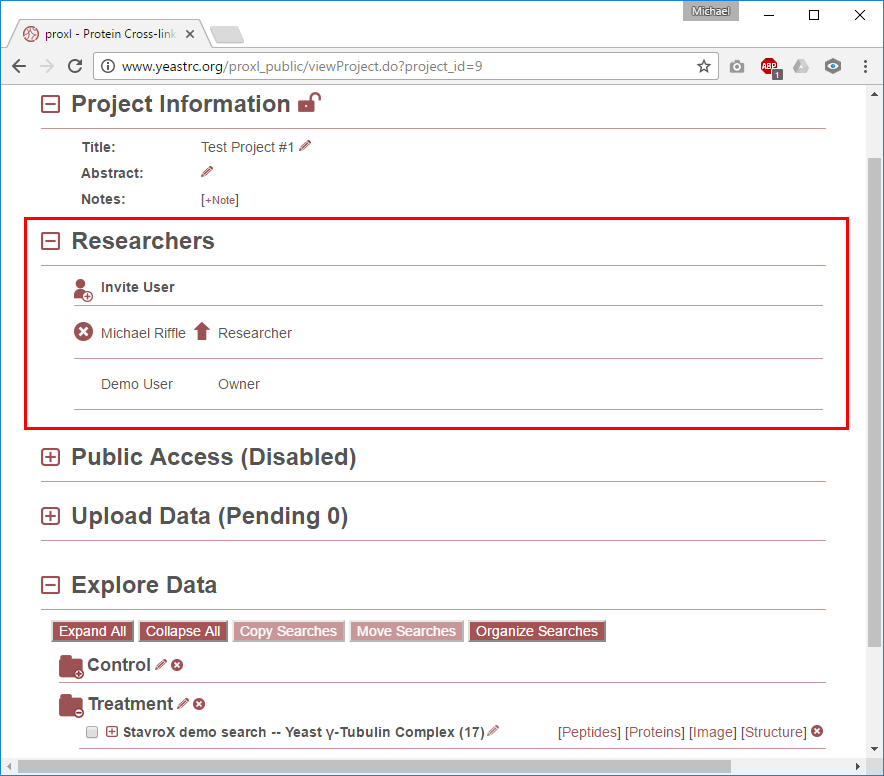
This section defaults to not being expanded, click the [+] icon next to this section title to expand it. This section lists the users associated with this project. Some important notes about the researchers section:
- Only users listed here may access the project overview page or the project’s data. (Except if public access is enabled, see below.)
- The “Researchers” section of this page is not visible to public users.
- Any user may invite other users to the project.
Invite User¶
To invite a user to the project, click the “Invite User” text or associated icon at the top of the user list. This will open the following dialogue.
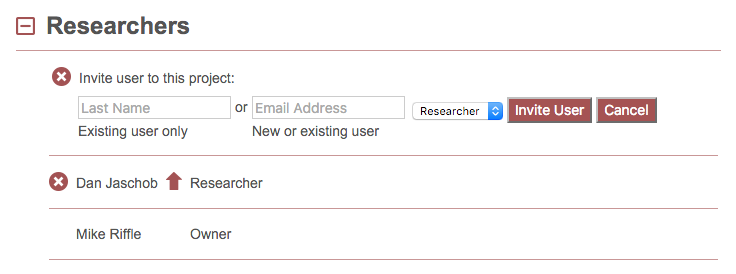
For Existing Users¶
To invite an existing user to this project, type their last name or email address into the respective text box. If they are found in the database, their name will appear as you type:
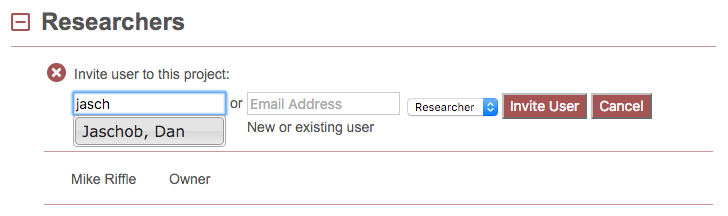
Clicking on the name produces:
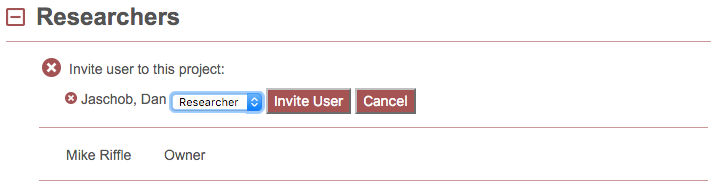
Here you may designate their level of access (if you are a project owner) and click “Invite User” to invite that user to the project. The user will immediately have access to the project, and this project will appear in their list of projects. Alternatively, click “Cancel” to cancel the process.
For New Users¶
To invite a new user to proxl and provide them access to this project, type in their email address and click “Invite User”. They will receive an email with a link for creating an account in proxl and this project will appear in their project list. New users invited to the project will appear as:
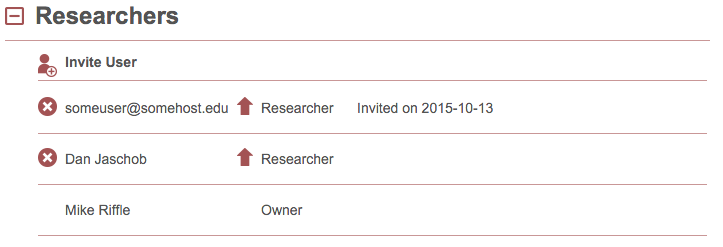
Once they have created an account, their name will appear in the user list instead of an email address, and the “Invited on…” text will no longer be present.
Remove User¶
Click the red (X) icon to the left of a listed user to remove that user from this project. That user will immediately lose access to the project and its associated data. This will not remove existing users, only revoke their access to this project. For invited email addresses, this will invalidate the access code included in the invitation email and they will not be able to use it to create a proxl account.
Promote or Demote User¶
Clicking the up or down arrow next to the access level of a user will either promote that user to an owner or demote that user to a researcher. Owners have complete complete access to a project, including the ability to lock or unlock it, enable or disable public access, promote or demote other users, define default views of data, or delete data.
Public Access¶
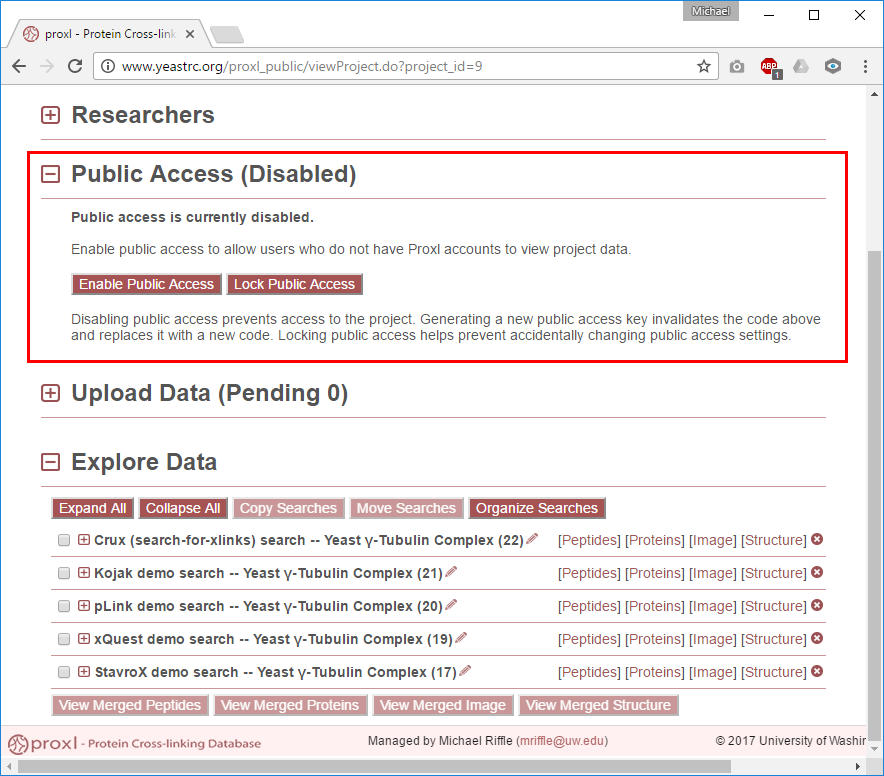
This section defaults to not being expanded, click the [+] icon next to this section title to expand it. This section controls whether or not public access is enabled and, if it is, whether or not a public access code is required to access the project and its data. Some important notes about the public access section:
- Only project owners may change public access settings.
- The “Public Access” section of this page is not visible to public users.
Enable or Disable Public Access¶
Enabling public access allows access to the data without requiring that users have proxl accounts. Clicking “Enable Public Access” enables public access and changes the display of this section to indicate that public access is enabled.
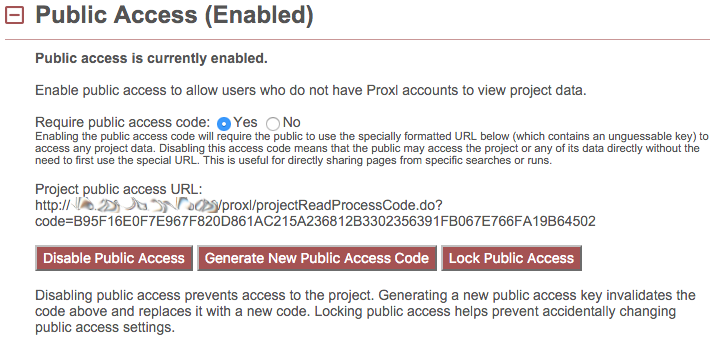
Public Access Code¶
By default, public access is enabled in a way that requires a specially-formatted URL that contains an unguessable public access code. This URL is listed here as “Project public access URL.” This exact URL must be used to access the project before the user may access any of the data. This is useful for semi-private sharing of data with select users (or reviewers) without making the data completely publicly accessible.
The requirement for the public access code may be removed by clicking “No” next to “Require public access code.” If “No” is selected, URLs for the project or any of the data pages may be directly shared without the need of the user to first use the public access code. This is useful for truly public sharing of the data, such as in the case of publication.
The “Generate New Public Access Code” button will generate and replace the current unguessable public access code with a new code. This will revoke access to users that have used the previous code.
Lock Public Access¶
Clicking the “Lock Public Access” button makes it impossible to change public access code settings without first clicking “Unlock Public Access.” This is meant to prevent accidental disabling of public access or generation of new public access codes, which would revoke previously-granted access to the public, colleagues or reviewers.
Upload Data¶
We have set up a separate page describing uploading data. Please see Upload Data.
Explore Data¶
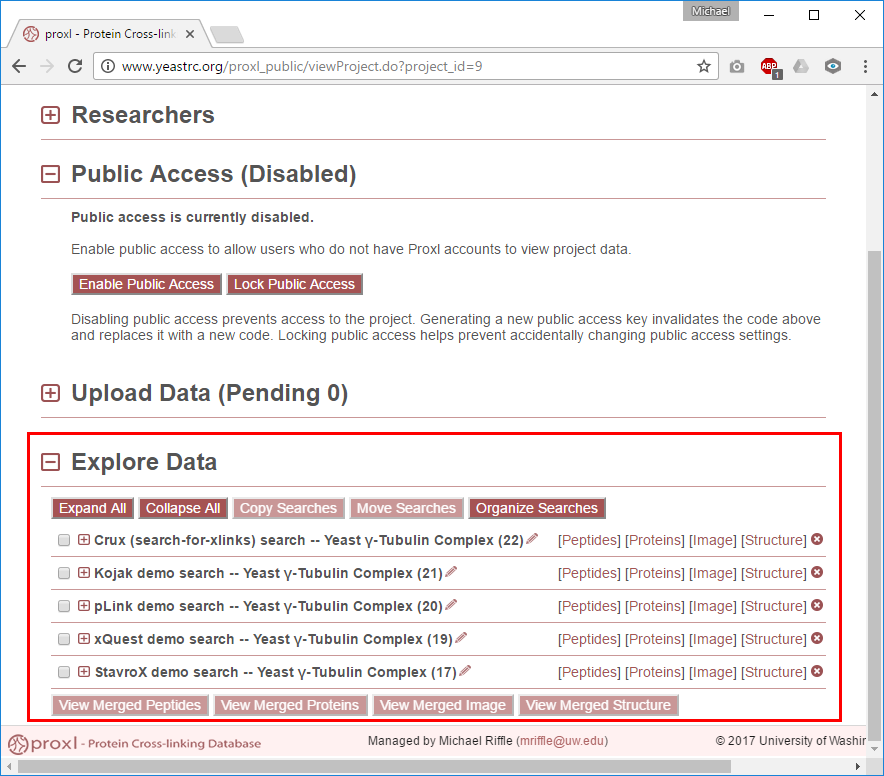
This section lists each of the searches associated with this project. A “search” in this context are all the data resulting from running a software pipeline (e.g., Kojak or xQuest) against spectra data (e.g., a mzML file). Project researchers may change the name of these searches by clicking the pencil icon to the right of the current search name. To the right of the search name the search ID number is listed in parentheses as a standard way to refer to specific searches.
View Search Information¶
To view information about a search, click the [+] icon to the left of the search name. This will display the following information. (Alternatively, click the “Expand All” button at the top of the to see all information about all searches.)
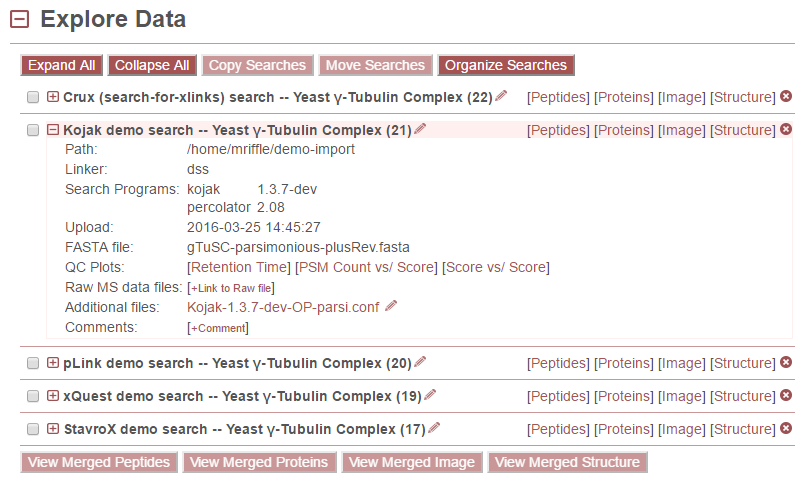
Search information includes the following information for each search:
- Path - The path the data were in when imported into proxl.
- Linker - the crosslinker(s) used in this experiment
- Upload - the date the data were uploaded to proxl
- QC Plots - links to a retention time QC plot. See below.
- Raw MS data files - Links to RAW files that contain the raw machine output for this experiment. Project owners may add URL links to RAW files by clicking [+Link to Raw file].
- Additional files - Links to additional files associated with this search, such as the configuration or parameters files for the respective search program.
- Comments - Lists the comments that have been added to the search. Comments may be deleted by clicking the red (X) to the left of the comment, or edited by clicking the pencil icon to the right.
Retention Time QC Plot¶
Clicking the “[Retention Time]” link next to “QC Plots:” produces the retention time QC plot:
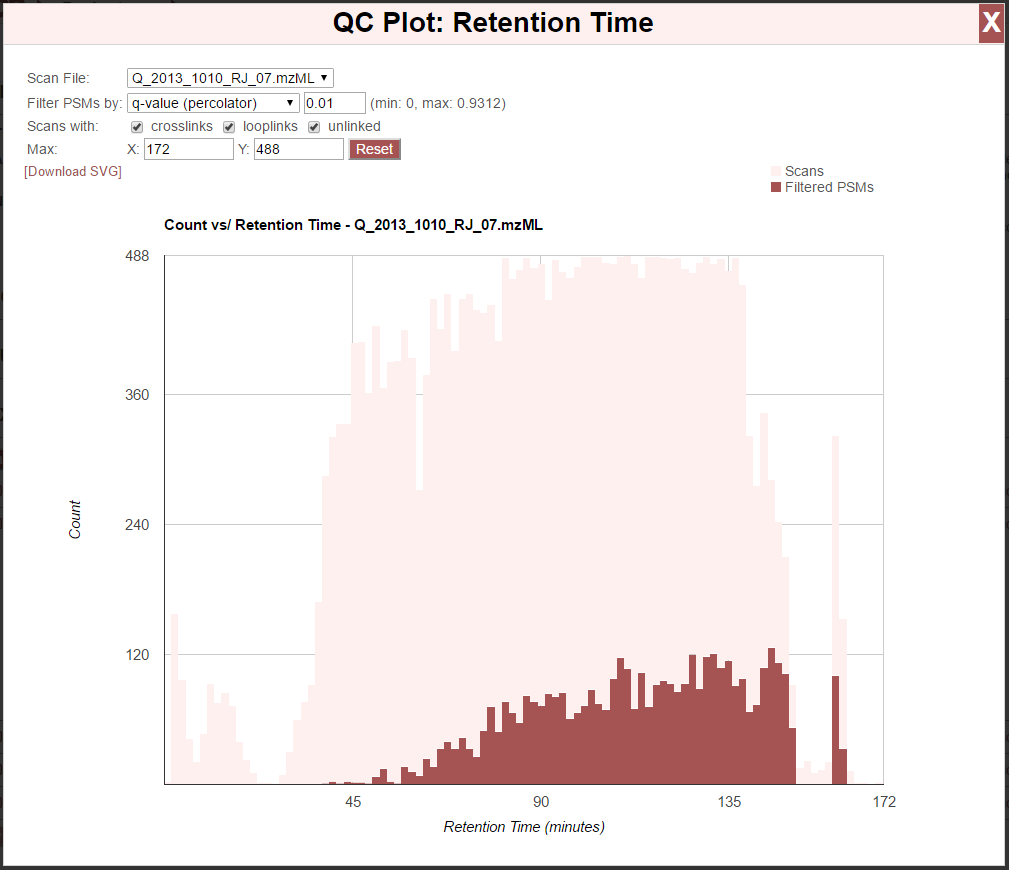
This is a histogram showing the number of MS2 scans taken versus retention time. The pink bars show all scans, and the dark red bars show the number of those scans that meet our filtering criteria at the top of the plot–or, “Filtered PSMs”. To close the chart, click the “X” in the top right of the window, or anywhere in the browser outside the chart window.
- The chart options are:
- Scan File - If multiple spectral files were searched, each will be listed here. The data in the chart reflect the selected spectral file.
- Filter PSMs by - Select the score type and cutoff value for that score to be used to plot the number of PSMs meeting those filtering criteria (red bars). The minimum and maximum values for the selected score type are given in parentheses.
- Scans with - Counts for “Filtered PSMs” will only include scans that resulted in a PSM where the peptide was of a type that is checked here. E.g., if only “crosslinks” is checked, only scans that resulted in crosslinked peptides will be used to for “Fitlered PSMs” counts. If “looplinks” and “crosslinks” are checked, only PSMs resulting in crosslinked or looplinked peptides will be counted.
- Max - Values entered here will be the maximum value on the X or Y axis–used for rescaling the chart.
Cumulative PSM Count vs/ score QC plot¶
Clicking the “[PSM Count vs/ Score]” link next to “QC Plots:” produces a plot of cumulative PSM count vs/ a chosen score type:
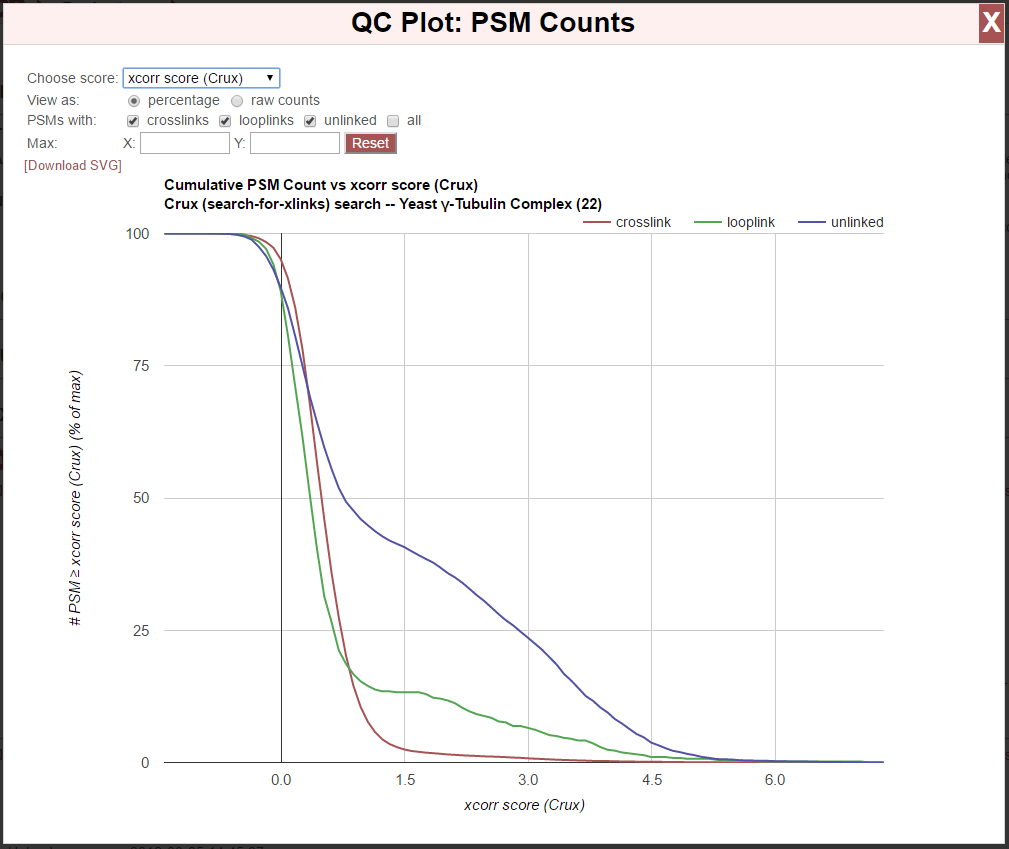
Possible values for the chosen score type are presented along the x-axis. Scores for which lower values are more significant, the y-value represents the number of PSMs with the value on the x-axis or lower. Score for which higher values are more significant, the y-value represents the number of PSMs with the value on the x-axis or higher.
Each class of PSM is presented as a separate line: crosslinks, looplinks, and unlinked. A line for all PSM types combined may optionally be displayed by checking “all” in the “PSMs with:” options.
The “Choose score:” option allows choosing which score from the search is used to generate the plot.
The “View as:” option allows switching between raw counts (default) and percentage. The “raw counts” option uses the raw PSM counts for the respective types. The percentage option displays the PSM counts as the percentage the total PSMs of the respective type, and so applies the same scale to all lines, which will always move from 0 to 100 in the plot.
The “PSMs with:” options toggle the visibility of the lines corresponding to the respective class of PSM.
The “Max:” options allow for a user-supplied maximum for the X- and Y-axes. “Reset” clears the user-supplied maximum values.
PSM Score Vs Score QC Plot¶
Clicking the “[Score vs/ Score]” link next to “QC Plots:” produces a plot of any PSM-level score vs/ any PSM-level score:
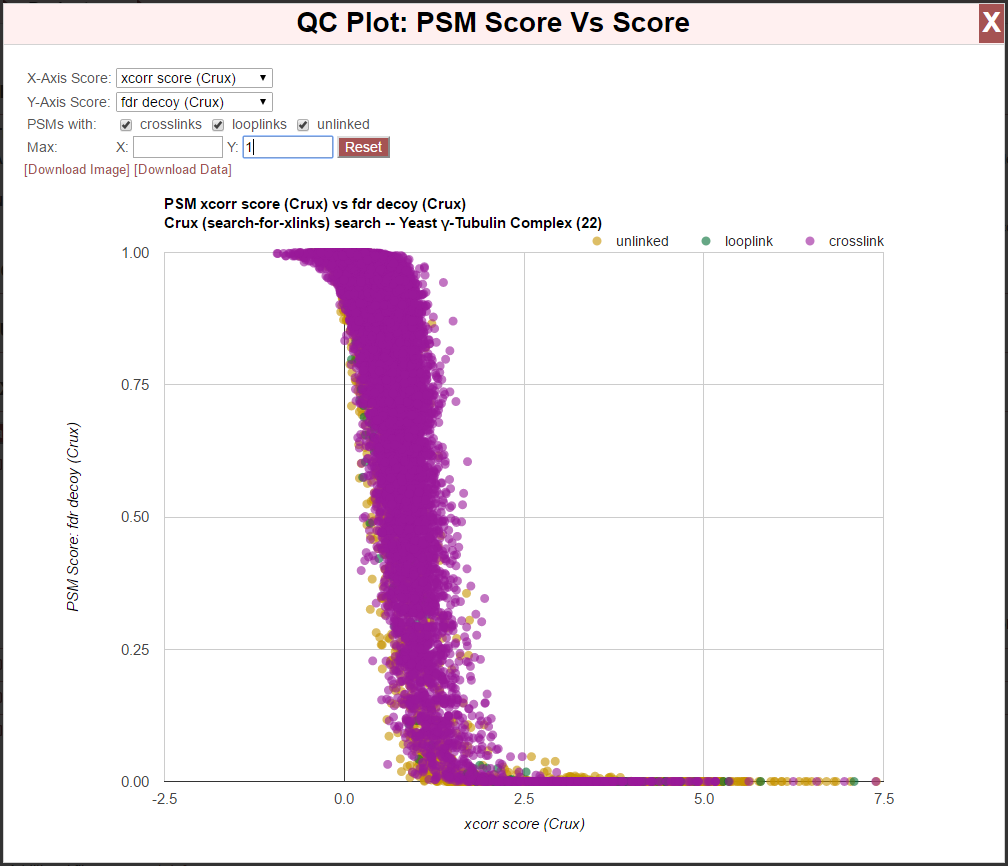
This is a scatter plot showing how PSM-level scores correlate with other PSM-level scores. For example, the above image shows calculated false discover rates (FDR) for PSMs vs/ the underling Xcorr score calculated by Crux. This can be used to discover unexpected relationships between scores, or to understand the effects of post processing statistical tools.
- The chart options are:
- X-Axis Score - The score to use for the x-axis.
- Y-Axis Score - The score to use for the y-axis.
- PSMs with - Select the type of PSMs to plot (cross-links, loop-links, and/or unlinked).
- Max - Values entered here will be the maximum value on the X or Y axis–used for rescaling the chart.
View Data¶
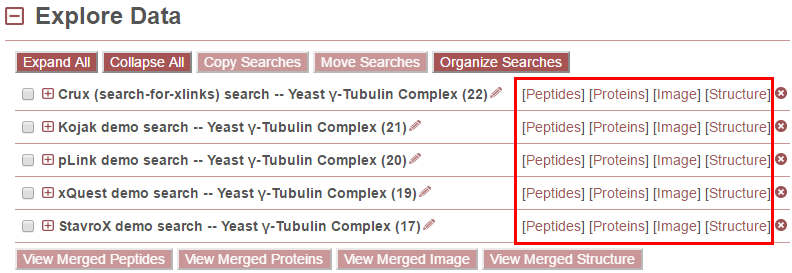
Use the links to the right of the search names (in red box above) to view the data. There are four views currently available:
- Peptides - Provides a table view of the identified peptides and associated data. See Peptides View Page for more details.
- Proteins - Provides a table view of the crosslinks and looplinks at the protein level. See Proteins View Page for more details.
- Image - Provides a graphical view of the data where proteins are represented as proportionately-sized bars that are annotated with link locations within and between proteins in the context of sequence annotation data. See Image View Page (Protein Bar View) for more details.
- Structure - Provides a view of crosslinking data on 3D protein structures, including an interface for uploading PDB files and mapping sequences from the FASTA file onto those PDB files using pairwise sequence alignment. See Structure View Page for more details.
View Merged Data¶
Proxl allows for comparing and contrasting multiple searches, even if those searches were analyzed with different software pipelines. Proxl refers to this as merging data. To merge data from multiple searches, click the check boxes to the left of the searches of interest and click either “View Merged Peptides”, “View Merged Proteins”, “View Merged Image”, or “View Merged Structure.”
For more information please see: Merged Peptides View Page, Merged Protein View Page, Image View Page (Protein Bar View), and Structure View Page.
Copy Searches¶
Project owners may copy searches to a different project (where they must also be an owner). To copy data, check the checkbox to the left of the search name(s) you wish to copy and click the “Copy Searches” button above the search list to see the following dialogue:
Projects to which you have permission to copy data are listed. Click on the project name to which the searches should be copied. Copied searches are treated independently with regards to the search name, comments, and other search metadata. For example, adding a comment or changing the name of the search in the new project will not affect the original search’s name or comments.
Move Searches¶
Project owners may move searches to a different project (where they must also be an owner). To move data, check the checkbox to the left of the search name(s) you wish to copy and click the “Move Searches” button above the search list to see the following dialogue:
Projects to which you have permission to copy data are listed. Click on the project name to which the searches should be copied. Moving a search will remove it from the current project and place it in the new project.
Organize Searches¶
The “Organize Searches” button opens an interface for rearranging the search list and creating and placing searches inside of “folders” for improved organization when the project contains many searches.
Clicking the “Organize Searches” button opens the following dialog:
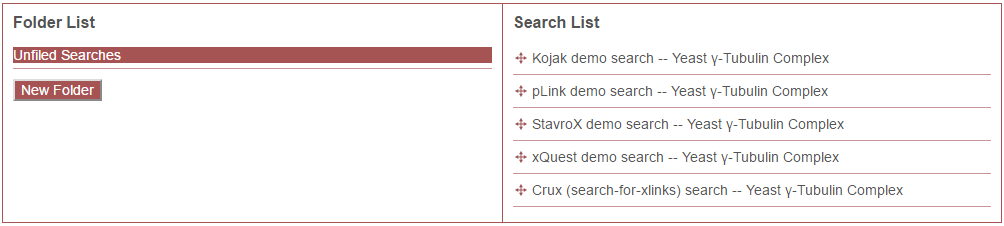
The right-hand panel is labeled “Search List”. This lists the searches contained in the currently-selected folder. To re-arrange the order in which searches are listed, click and drag the search to the desired order in the list. To see and re-arrange searches in another folder, click on the name of the folder in the left-hand panel.
The left panel is labeled “Folder List”. This is a list of the folders that have been created for organizing searches. Note: Searches not in any folder are listed here under a special folder named “Unfiled Searches”. Any searches in “Unfiled Searches” will not be placed into a folder when shown to users in the web application.
To create a new folder, click the “New Folder” button, type in the name of the new folder, and click “Add Folder.” In the following example, two folders have been created, “Control” and “Treatment.”
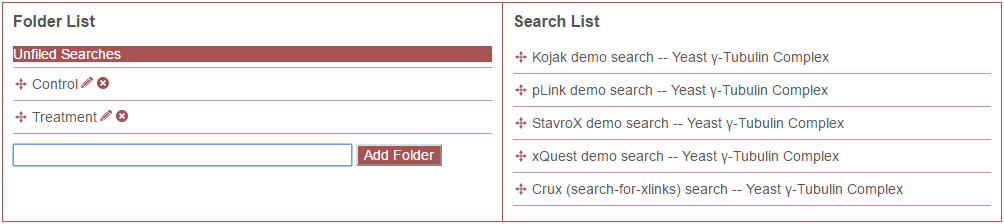
Folders may be deleted by clicking the small red “X” icon next to the folder name. Folder names may be edited by clicking the pencil icon. And folders may be re-arranged by clicking and dragging them to the desired position.
To place a search in a folder, first click on the folder name that currently contains the search (remember, searches not in folders are under “Unfiled Searches”). Then click the name of the search in the right-hand panel and drag it onto the row containing the folder in the left-hand panel and release the mouse button. This will “drop” that search into that folder.
When done, click the “Done Organizing Searches” button above the folder list to return to the normal interface.
In the following example, the StavroX and xQuest demo searches were added to the “Treatment” folder. The Crux demo search was left unfiled. And the user has clicked on the “Treatment” folder to view the searches listed under it.
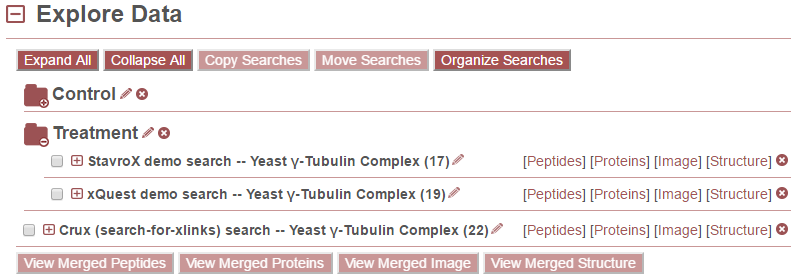
Delete Searches¶
Project owners may delete searches by clicking the red (X) icon to the right of the view data links. This will remove all data associated with that search from the database–use with care. Note that if this search was copied to another project, that copy search will not be deleted.